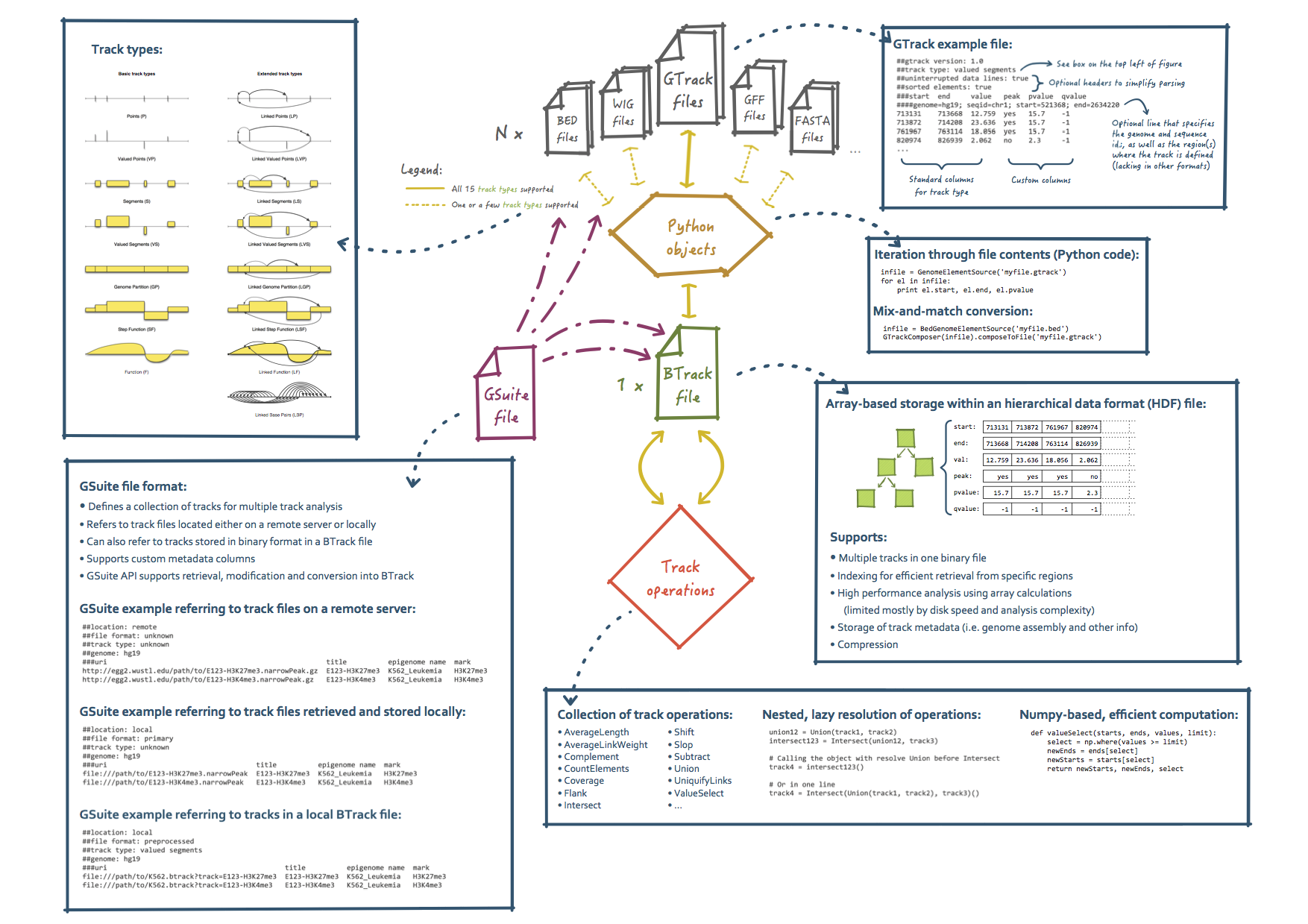
GTrack, BTrack and GSuite are file formats designed to handle genomic track data of heterogeneous types. The file formats are designed to complement each other and work jointly as a complete ecosystem for representation and analysis of most types of data that can be located along a reference genome.
GTrack[1] is a tabular format that was developed as part of the Genomic HyperBrowser[2,3] system to provide a uniform representation of most types of genomic datasets. GTrack is able to replace common formats such as WIG, GFF, BED, FASTA, in addition to represent chromatin capture datasets, such as Hi-C and ChIA-PET.
The BTrack format supports the same variety of informational content as GTrack, but in a binary form. BTrack is unique in supporting a collection of multiple tracks stored together in one HDF5-based binary file, while still supporting a high level of efficiency. It is currently implemented as a prototype and not finalized.
The GSuite[4] format is a unique tabular format that binds together the whole chain of multi-track analysis, from search and retrieval of genomic tracks, through intermediate processing, to analysis.
A Python package supporting parsing, conversion and operations is available from https://github.com/gtrack/gtrackcore.
- Gundersen, Sveinung, et al. "Identifying elemental genomic track types and representing them uniformly." BMC Bioinformatics 12.1 (2011): 1. DOI: 10.1186/1471-2105-12-494
- Sandve, Geir K., et al. "The Genomic HyperBrowser: inferential genomics at the sequence level." Genome biology 11.12 (2010): 1-12. DOI: 10.1186/gb-2010-11-12-r121
- Sandve, Geir K., et al. "The Genomic HyperBrowser: an analysis web server for genome-scale data." Nucleic acids research 41.W1 (2013): W133-W141. DOI: 10.1093/nar/gkt342
- Simovski, Boris, et al. "GSuite HyperBrowser: integrative analysis of dataset collections across the genome and epigenome." GigaScience 6.7 (2017): 1-12. DOI: 10.1093/gigascience/gix032