AICSImageIO bindings for napari
- Supports reading metadata and imaging data for:
CZIOME-TIFFTIFF- Any formats supported by aicsimageio
- Any additional format supported by imageio
Stable Release: pip install napari-aicsimageio
Development Head: pip install git+https://github.com/AllenCellModeling/napari-aicsimageio.git
There are two variants of this plugin that are added during installation:
aicsimageio-in-memory, which reads an image fully into memoryaicsimageio-out-of-memory, which delays reading ZYX chunks until required. This allows for incredible large files to be read and displayed.
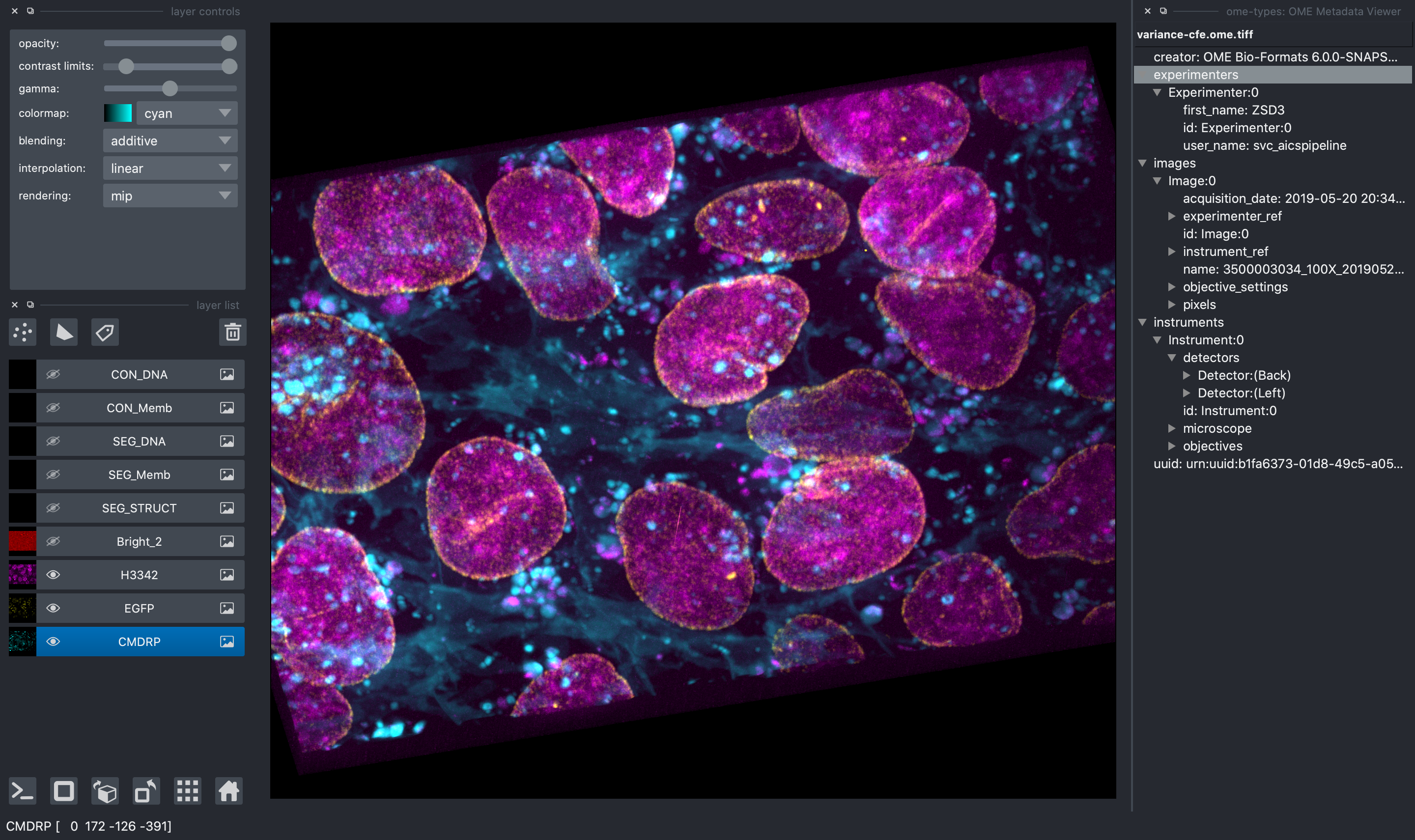
All image file formats supported by aicsimageio will be read and all raw data will be available in the napari viewer.
In addition, when reading an OME-TIFF, you can view all OME metadata directly in the
napari viewer thanks to ome-types.
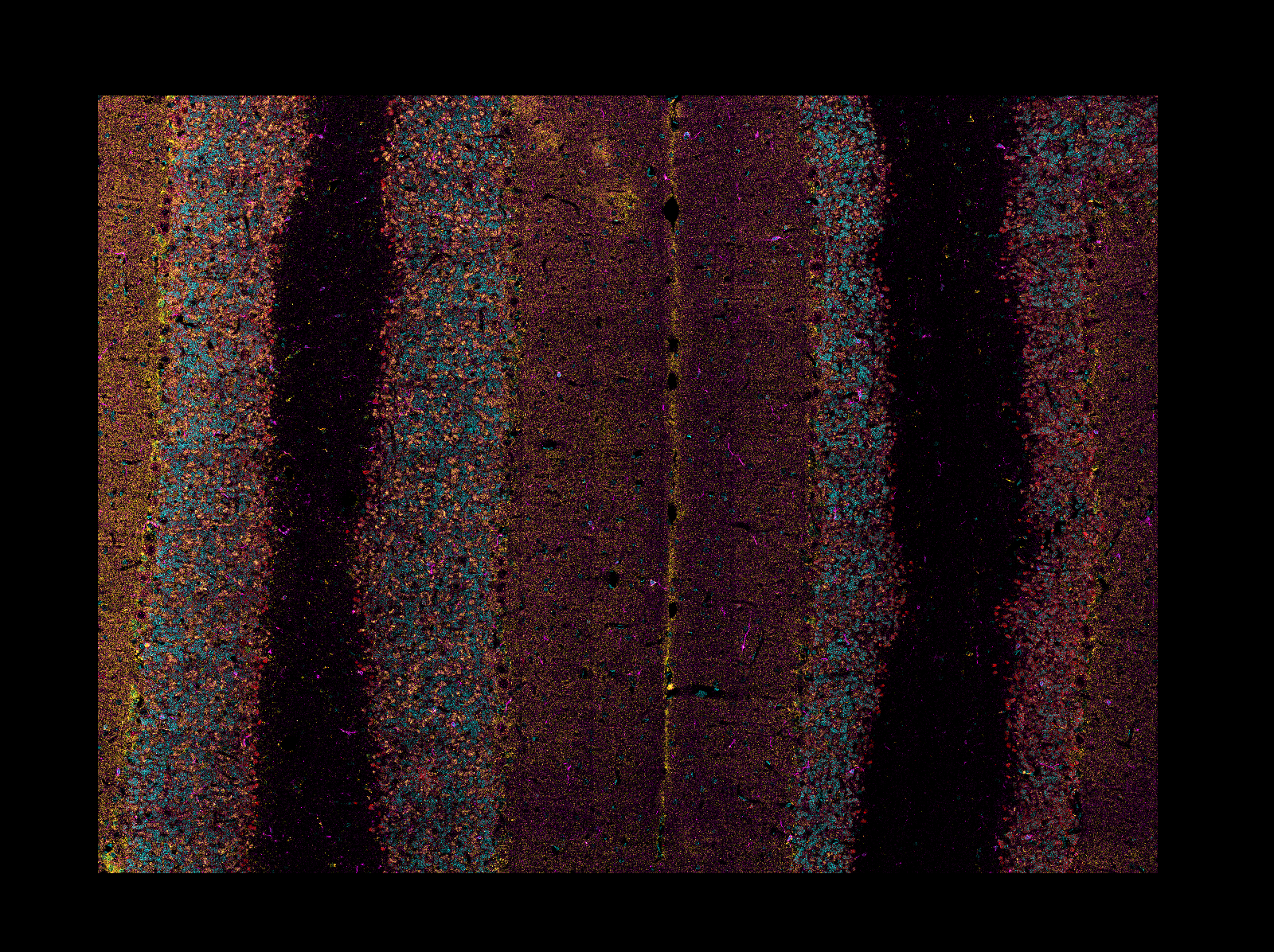
When reading CZI or LIF images, if the image is a mosaic tiled image, napari-aicsimageio
will return the reconstructed image:
Experimental
When reading a multi-scene file, a widget will be added to the napari viewer to manage scene selection (clearing the viewer each time you change scene or adding the scene content to the viewer) and a list of all scenes in the file.
See CONTRIBUTING.md for information related to developing the code.
For additional file format support, contributed directly to
AICSImageIO.
New file format support will become directly available in this
plugin on new aicsimageio releases.
If you find aicsimageio (or napari-aicsimageio) useful, please cite as:
AICSImageIO Contributors (2021). AICSImageIO: Image Reading, Metadata Conversion, and Image Writing for Microscopy Images in Pure Python [Computer software]. GitHub. https://github.com/AllenCellModeling/aicsimageio
Free software: BSD-3-Clause