ofxTSNE is an addon for openframeworks which wraps the t-SNE (t-Distributed Stochastic Neighbor Embedding) algorithm by Laurens van der Maaten.
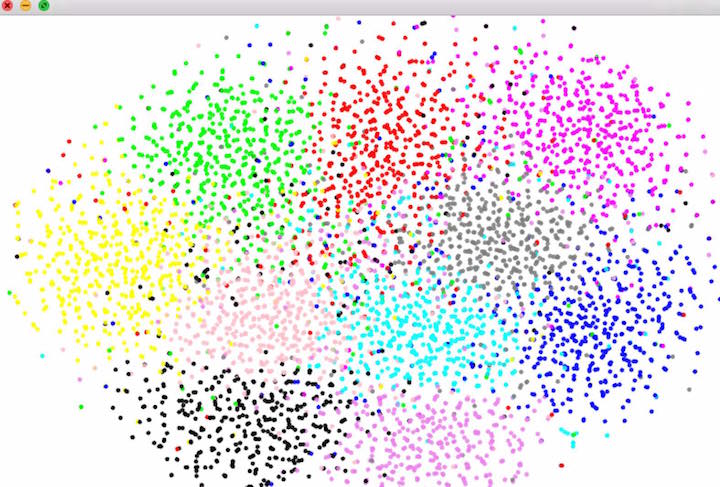
t-SNE is a technique for reducing the dimensionality of large, high-dimension datasets, typically to 2 or 3 dimensions. It has a similar function to Principal Component Analysis (see ofxPCA) which reduces a dataset's dimensionality by reorienting it along its principal axes, but differs in that it tends to better preserve point-wise distances, making it more suitable for visualization of high-dimensional data.
ofxTSNE is very simple to run, containing only one function. The harder part is getting data.
example demonstrates how to use ofxTSNE by constructing a toy 100-dim dataset. It contains comments explaining what the parameters do and how to set them.
clever hack: try setting D=3 and instead of making points clustered around 10 centers, make the points random 3d points and map the point's color linearly from its 3d position.
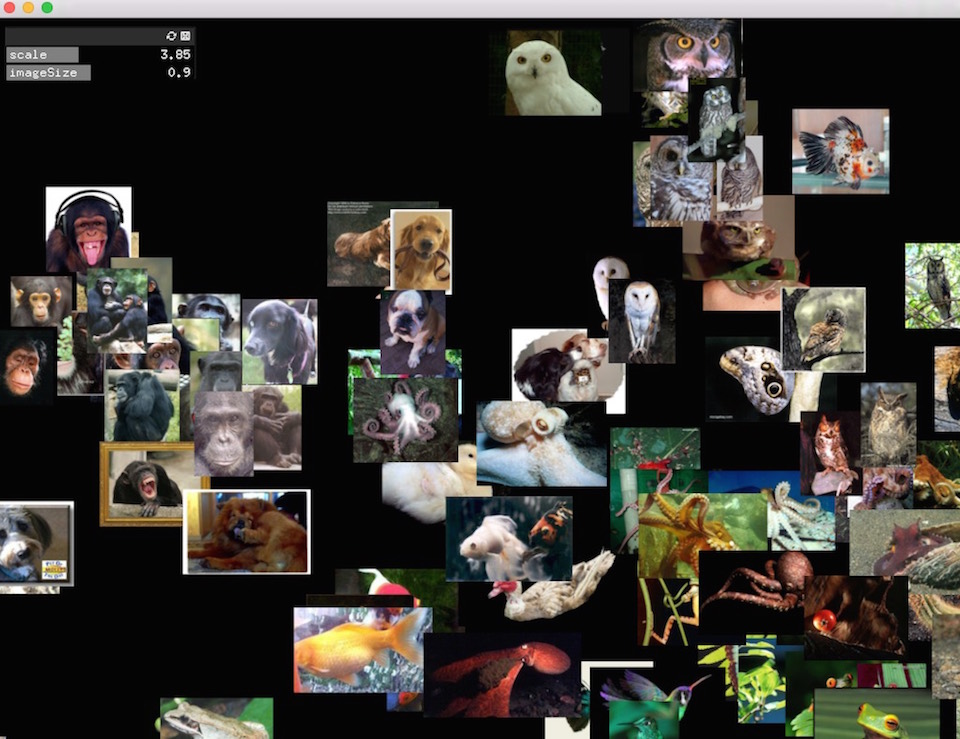
example-images applies t-SNE to a directory of images. It uses ofxCcv to encode each image as a compact (4096-dim) feature vector derived from a trained convolutional neural network. The resulting representation captures high-level similarities among images, enabling ofxTSNE to group them effectively according more to content (e.g. images of cats get clustered together), relatively invariant to changes in color, lighting, position, etc.
To run this example, you need to take a few extra steps.
-
Get ofxCcv
-
run the setup_ccv script to download the trained convnet.
sh setup_ccv.sh
-
Then you need to populate a folder called 'images' inside your data folder. Be careful to use small-sized images because the entire directory will be loaded into memory. I've provided a script which downloads 20 images each from 31 categories in Caltech-256. If you'd like to download those, run:
python download_images.py
Or if you want to download a set of animals from the same source, open download_images.py and change the line categories = categories_random to categories = categories_animals.
You can easily find an optimal 2d grid which preserves your t-SNE clusters using a 2d assignment algorithm. @Quasimondo's RasterFairy or Kyle McDonald's ofxAssignment will work, or something similar. Applying RasterFairy to a t-SNE of the animals subset from Caltech-256 (see download_images.py), you get a result that looks like the following: