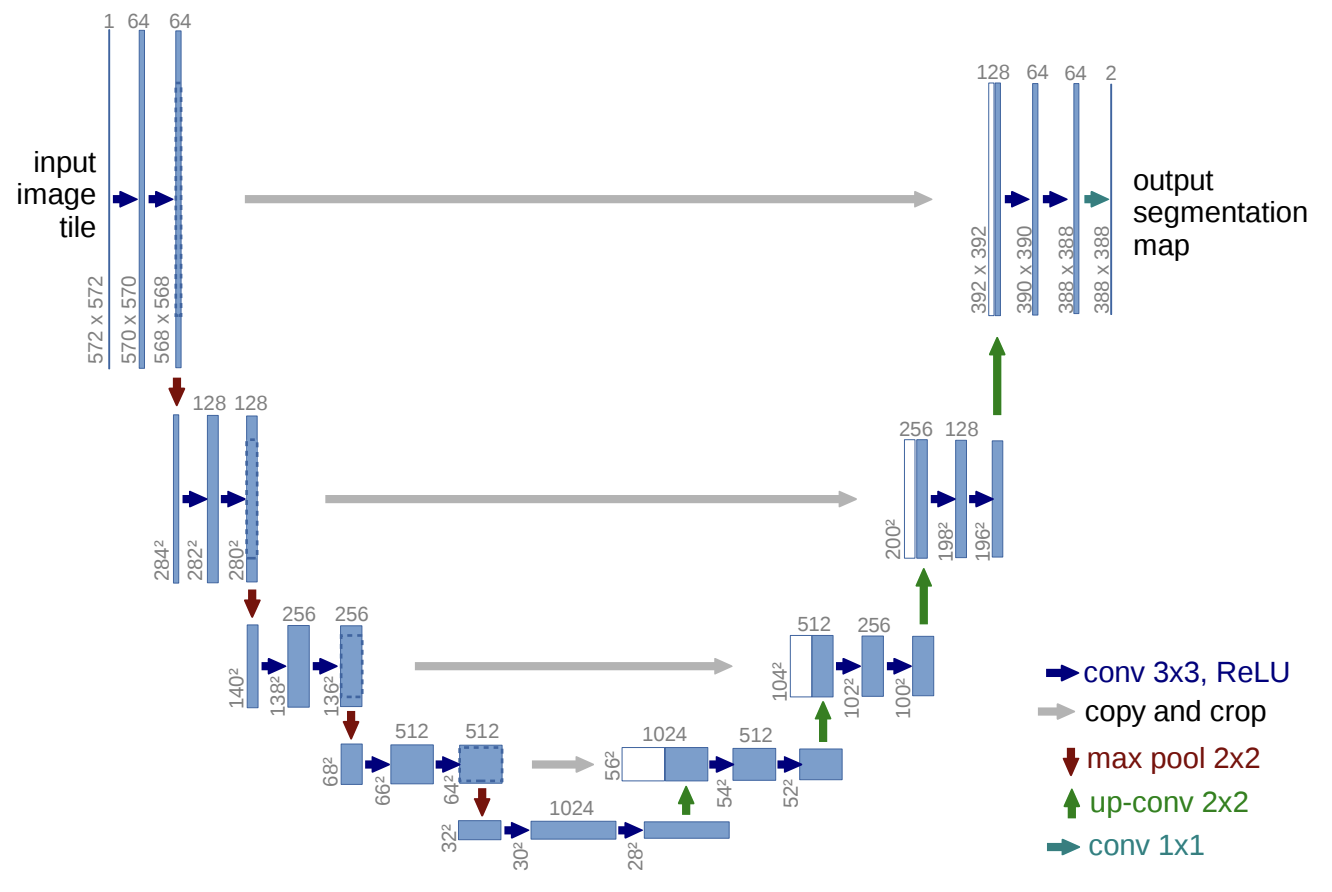
U-Net is a convolutional neural network architecture, originally developed for biomedical image segmentation. It features a symmetric, U-shaped structure with a contracting path to capture context and a symmetric expanding path for precise localization, enabling it to effectively segment images even with limited training data. U-Net is widely used in medical image analysis and other segmentation tasks.
U-Net architecture in the original paper:
.
├── LICENSE
├── README.md
├── config.yaml
├── data.py
├── model
│ ├── Trainer.py
│ └── UNet.py
├── model-params
├── requirements.txt
├── result
└── train_model.py-
Install Dependencies
Begin by installing the required packages using
pippip install -r requirements.txt
-
Configure Your Model
Next, tailor the
config.yamlfile to your specific requirementsmodel-conf: num_classes: 2 # Define the number of classes train-conf: w_c: True # Include class frequency in the loss function pixel weight w_d: False # Include pixel border distance in the loss function pixel weight (TODO) num_epochs: 20 # Set the number of training epochs batch_size: 8 # Specify the batch size lr: 0.0001 # Set the learning rate
-
Prepare Your Dataset
In
data.py, complete the dataset preparation stage:def load_train_dataset(): print('Loading training dataset...') # TODO return [] def load_val_dataset(): print('Loading validation dataset...') # TODO return [] def load_test_dataset(): print("Loading test dataset...") # TODO return []
Ensure that each function returns a list of tuples
(img, mask). In this context,imgshould be a three-channel tensor, for instance, (3, 512, 512), andmaskshould be a one-channel integer tensor, such as (512, 512). The dimensions ofmaskmust correspond to those ofimg, with both the height and width being divisible by 32. Each element withinmaskrepresents the class of the pixel it corresponds to, ranging from0tonum_classes - 1. -
Initiate Training
Start the training process with the following command:
python train_model.py
Upon completion, the training and MIoU curves will be saved in the
resultdirectory. The model parameters for each epoch are stored inmodel-paramsas.pthfiles.
This revised version maintains the original technical accuracy while enhancing clarity and formality.
-
Complete
w_dgeneration task -
Complete road segmentation experiment
-
Complete object segmentation experiment on VOC2012 dataset